Epitope modeling and MHC project
Assistant researcher project in computational and medicinal chemistry (2015 - 2016)
University of Chile, Faculty of Chemical and Pharmaceutical Science
Peptides that come from the fractionation of pathogen proteins in intracellular lysosomes are recognised by the immune system by the binding of antigen-presenting proteins known as the histocompatibility complex (MHC) class I and II type. In order to design peptide vaccines I investigated the binding of short chain peptides candidates that could start an immune response as antigens and activate the T lymphocytes in order to initiate the immune response. The prediction of peptide fragments with potential affinity to bind to the MHC I or II could lead to the development of an optimal immune response to promote immunity in organisms such as humans or animals.
In this project I worked in collaboration with colleagues of University of Chile and a Chilean Veterinary company to find out potential peptide sequences to treat diseases in Salmon species such as *Salmon Salar* type. In order to predict peptide affinity in MHC-peptide complexes and to design antigens a molecular modelling protocol has been developed from studies involving techniques such as homology modelling of MHC I and II, prediction and design of peptide structures using *de novo* methods.
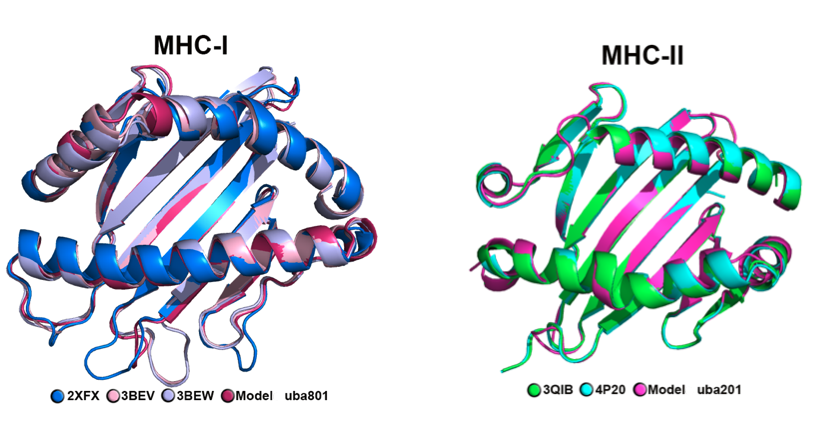
The *Salmon Salar* MHC models were obtained using homology modeling using *Modeller* and taking protein templates from Protein Data Bank. Different de novo design techniques based on Monte Carlo calculations available on Rosetta were studied to model peptides of 9 amino acid length. The affinity of best candidate peptide models with MHC was evaluated using Molecular Dynamics (GROMACS) and then calculated using the free energy calculation by the MMPBSA approach.