Hi! I am Montserrat
Explore my portfolio to see my work and learn more about my work and what I can offer you.

Explore my portfolio to see my work and learn more about my work and what I can offer you.

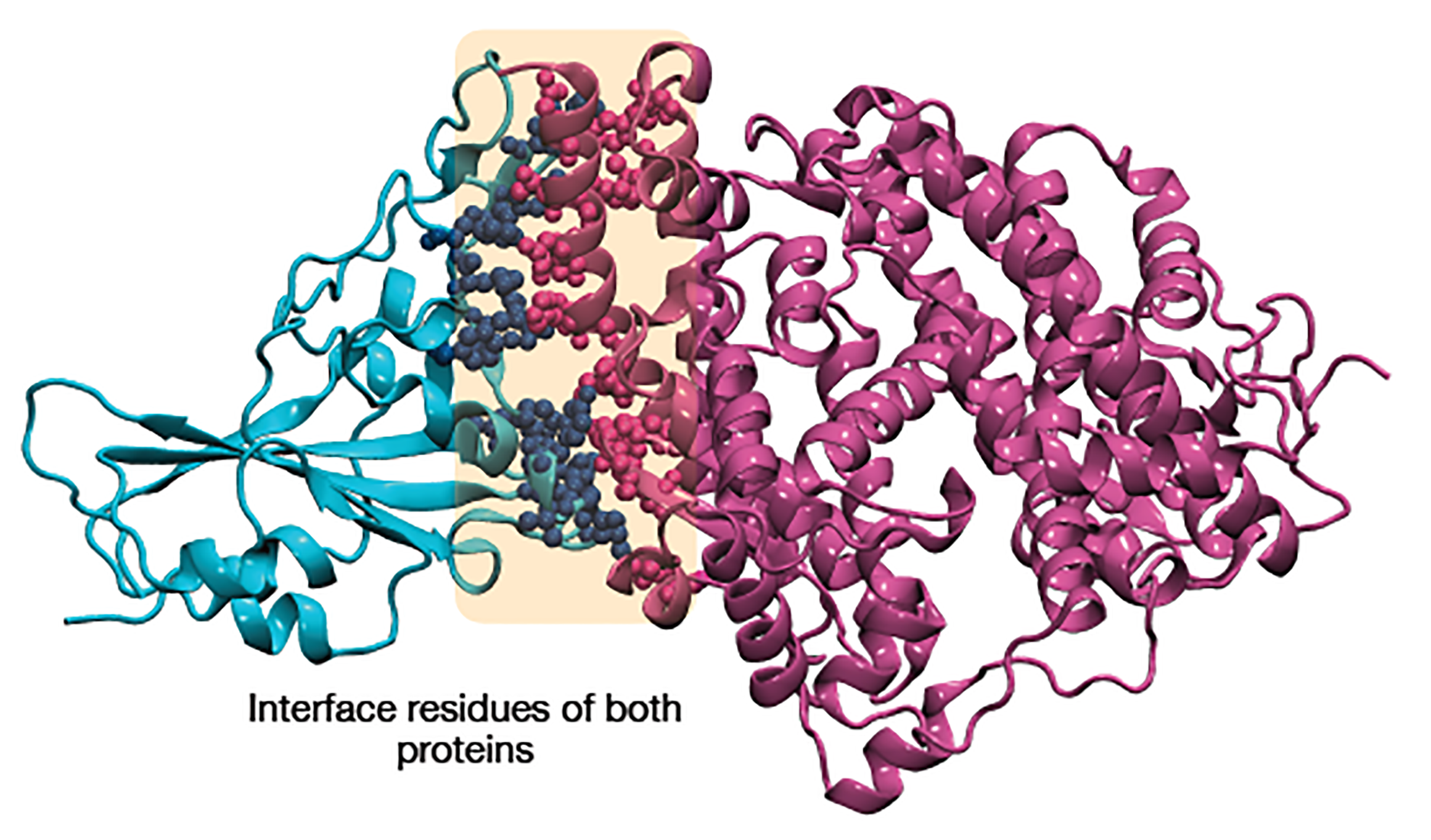
In this project I implemented python modules to analyse the interface of two protein interfaces to elucidate the key aminoacids interacting in the interface, aminoacid content and contact map
Skills: Python, OOP, Data Science, Structural Biology.
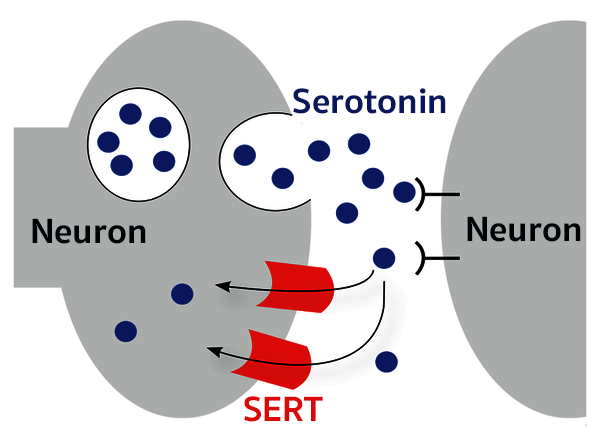
I investigated the structural patterns of Serotonin transporter ligands using Cheminformatic methods such as exploratort analysis, clustering methods and QSAR. The preliminary analysis is used for ML studies.
Skills: Python, Data Science, Cheminformatics, Machine Learning.

As a personal project, I performed exploratory analysis on MAO-A ligand dataset from CheMBL, and I used to perform Machine Learning (ML) and Deep Learning (DL) training.
Skills: Python, Data Science, Cheminformatics, Machine Learning.
I accomplished two projects showing the develop of complex simulations protocols and automating their execution. They include protocols for exploring the possible dihedral values on the backbone of the polymer and studying the possible conformations of sequence-controlled oligomers and making them accessible to non-computer scientists.
Skills: Python, noSQL, Molecular Dynamics, Monte Carlo, Density Functional Theory, HPC.
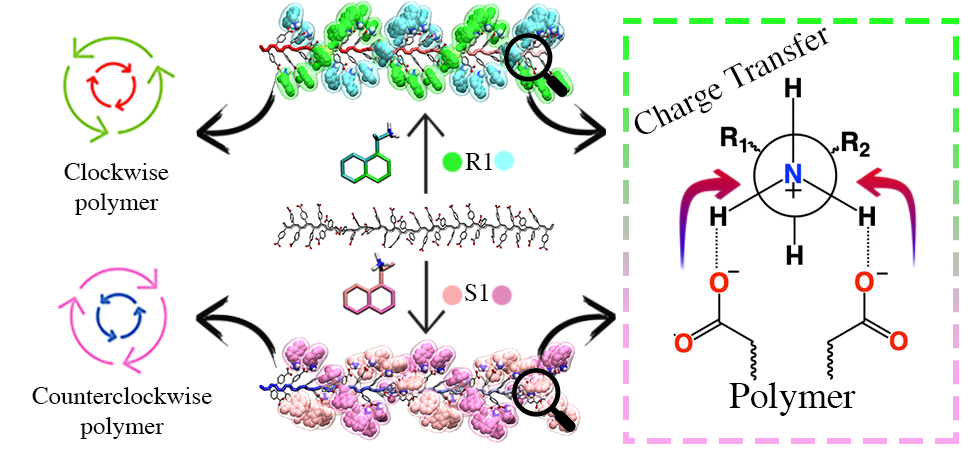
I studied the helical induction process of cis-transoid poly 4-(carboxyphenyl) acetylene by chiral amines combining computational Molecular Dynamics (MD) and Density Functional Theory (DFT) approaches.
Skills: Python, Molecular Dynamics, Density Functional Theory, HPC.
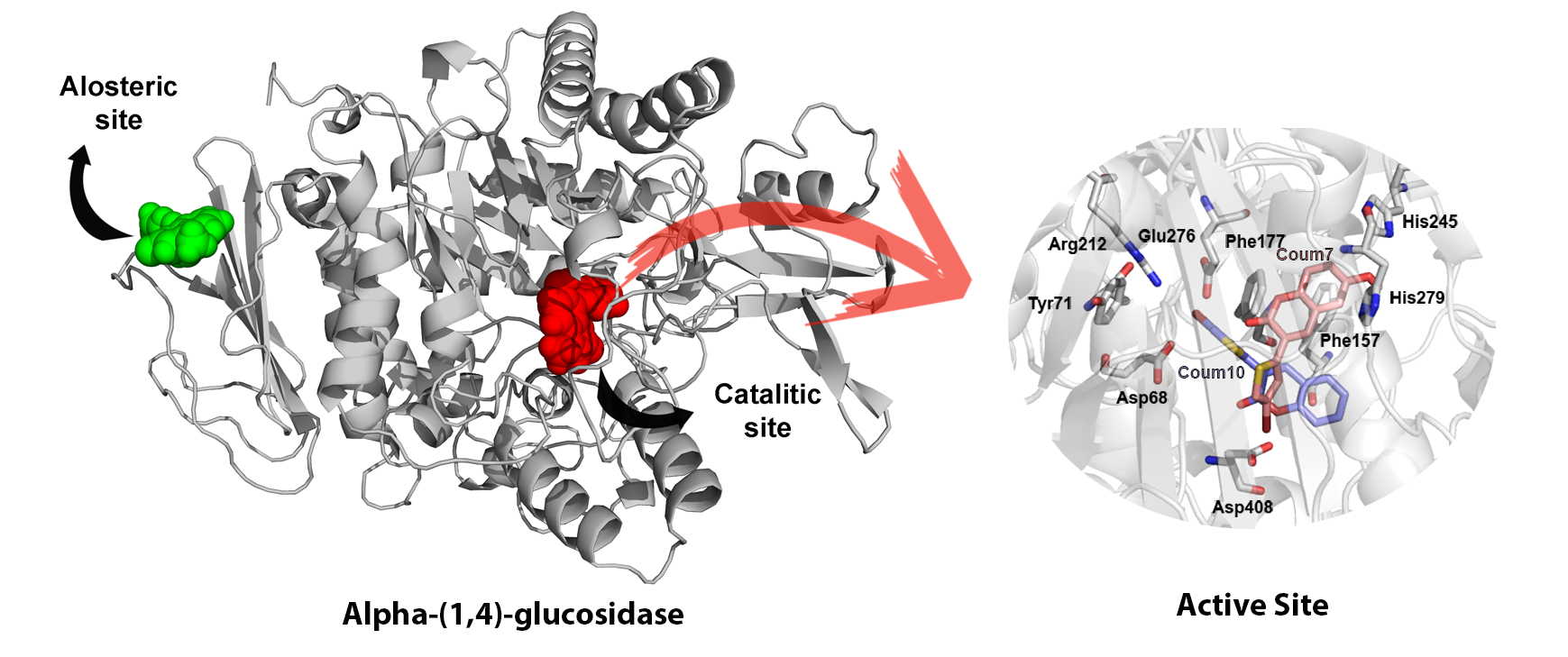
In collaboration with experimentalists, I studied the binding of potent coumarines on Diabetes Mellitus II protein targets (Publication in Current Topics in Medicinal Chemistry).
Skills: Medicinal Chemistry, BASH, Docking, Homoloy Modeling, HPC.
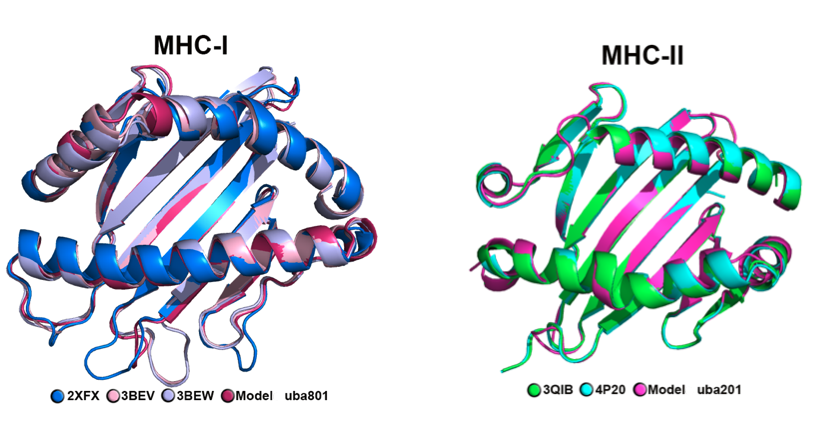
I implemented modeling protocols to perform de novo modeling to study epitopes that binds antigen-presenting proteins.
Skills: Medicinal Chemistry, Monte Carlo Simulations, Python, HPC.

I investigated how the path to active site of MAO-B could influence the activity on MAO as substrates and inhibitors using advanced Molecular Dynamic techniques.
Skills: Medicinal Chemistry, Molecular Dynamics, Docking, BASH, HPC.